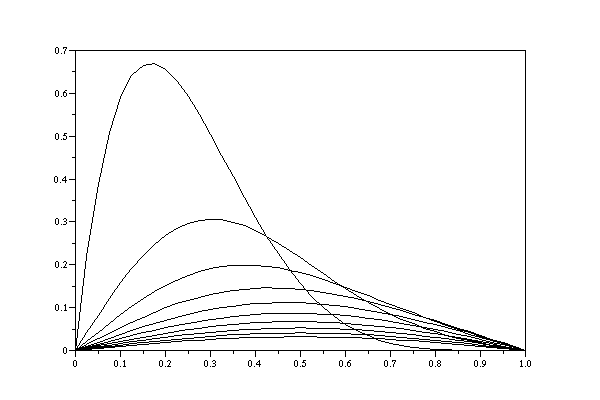
There are 6 functions below:
function a=u0(x) a=10*x.*(1-x).^5; endfunction
function x=tridiag(d,u,l,r)
// solve [ d1 u1 0 0 ] [ x1 ] = [ r1 ]
// [ l1 d2 u2 0 ] [ x2 ] [ r2 ]
// [ 0 l2 d3 u3 ] [ x3 ] [ r3 ]
// [ 0 0 l3 d4 .... ] [ x4 ] [ r4 ]
// [ : ] [ : ] [ : ]
ds=size(d,1)
us=size(u,1)
ls=size(l,1)
rs=size(r,1)
x=zeros(ds,1)
if (us==ds-1 & ls==ds-1 & rs==ds )
for i=2:ds
fact=l(i-1)/d(i-1)
l(i-1)=0;
d(i)=d(i)-fact*u(i-1)
r(i)=r(i)-fact*r(i-1)
end
x(ds)=r(ds)/d(ds)
for i=ds-1:-1:1
x(i)=(r(i)-u(i)*x(i+1))/d(i)
end
else
x=0 // this will give an error if input sizes don't match
end
endfunction
function a=step(u,nu) d=size(u,2); a=zeros(1,d); a(1)=0; a(d)=0; for i=2:d-1 a(i)=u(i)+nu*(u(i-1)-2*u(i)+u(i+1)); end endfunction
function s=parint(T,J,N)
x=(1/J)*[0:J]
u=u0(x)
t=0
s=0
nu=T*J*J/N
xbasc(0)
plot2d(x,u)
for i=1:N
u=step(u,nu);
t=t+nu/(J*J);
if (t-s>0.025)
plot2d(x,u)
s=t
end
end
endfunction
function unew=step2(u,nu,theta) dimu=size(u,1); D=zeros(dimu,1) U=zeros(dimu-1,1) L=zeros(dimu-1,1) R=zeros(dimu,1) D=D+(1+2*theta*nu) D(1)=1 D(dimu)=1 U=U-theta*nu U(1)=0 L=L-theta*nu L(dimu-1)=0 m1=1-2*(1-theta)*nu m2=(1-theta)*nu R(1)=0 R(dimu)=0 for i=2:dimu-1 R(i)=m1*u(i)+m2*(u(i-1)+u(i+1)) end unew=tridiag(D,U,L,R) endfunction
function s=parint2(T,J,N,theta)
// T=total time, J=steps in x-dirn, N=steps in t-dirn, theta
x=(1/J)*[0:J]'
u=u0(x)
t=0
s=0
nu=T*J*J/N
xbasc(0)
plot2d(x,u)
for i=1:N
u=step2(u,nu,theta);
t=t+T/N;
if (t-s>0.025)
plot2d(x,u)
s=t
end
end
endfunction

To see that the theta method does not require such large numbers of steps in the t direction, provided theta is at least 1/2, run parint2(1,40,80,0.4) (fails) and parint2(1,40,80,0.5), parint2(1,40,80,0.6) (both work fine). Note the vast improvement of speed over the explicit method!